Welcome to my Project Portfolio!
I'm Elliot Chan
Computational Biologist | Honours Biomedical Sciences Graduate
About Me
I've independently built end-to-end platforms that bridge biomedical research workflows and computation, from clinical variant analysis to drug optimization. My work combines bioinformatics, machine learning, and algorithmic innovation to solve complex problems in the computational side of healthcare. Outside programming I'm either training (Weights / Running / Muay Thai / Boxing), or reading. Actively looking for entry-level bioinformatics, computational biology, or ML roles.
GEM & PRISM: Screen, Repair, Interpret
GEM is an ML-enhanced framework designed to classify and repair gene
variants for reduced pathogenicity. Featuring a 500+ feature engineering
pipeline paired with my personal feature space optimization tool DataSift,
GEM has trained high-performance models with 90% ROC-AUC, 89.6% PR-AUC
and 82% Accuracy, and that's just from looking at the sequence and
chromosome number. ReGen, a system present in both GEM and PRISM,
is an iterative guided mutation algorithm capable of identifying gene
therapy targets when presented with a pathogenic gene variant.
PRISM extends GEM into a result-oriented worktool of its own.
Where GEM trains models and produces predictions, PRISM is
specialized towards producing and interpreting said results.
Possessing the same feature engineering pipeline and algorithms,
PRISM not only features a CLI system but also includes an
AI-interpretation layer, allowing for generation of biologically
grounded hypotheses based on result data as well as experimental
follow-up proposals. This allows users to drag + drop files into
PRISM then screen, repair, and interpret gene variants from the
CLI.
BlueTuna ><>
A multi-stage hyperparameter optimization engine for binary classifiers. Characterizes the parameter landscape before searching it.
Where traditional tuning libraries treat the search space as something to be continuously exploited, BlueTuna treats it as something to be understood beforehand. BlueTuna creatively addresses the task of hyperparameter optimization by breaking it down into 3 stages, each informing the next:
- [1] Parameter search space filtering via gradient-based region scoring, isolates regions most likely to contain performance optima
- [2] Custom perceptron training on Latin Hypercube-sampled data from filtered search space to better "see" the hyperparameter landscape
- [3] Fixed-weight gradient descent on hyperparameter values, guiding model performance towards optima
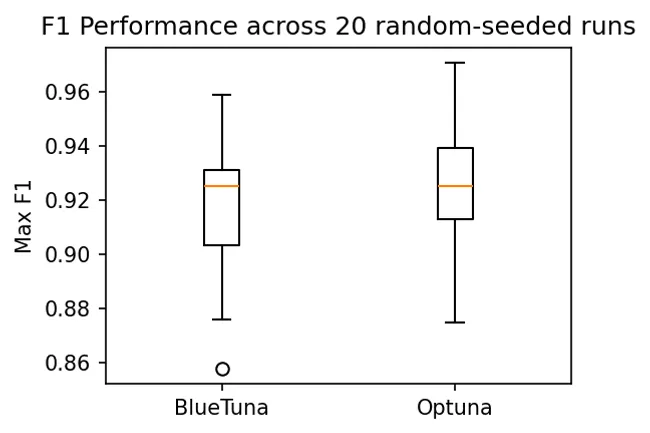
BlueTuna was able to achieve a competitive performance ceiling against Optuna (top-performing optimizer), beating it on 5/20 trials and competitive on 40%. Notably, its median performance slightly exceeded Optuna's, demonstrating its limitation to be consistency rather than performance ceiling.
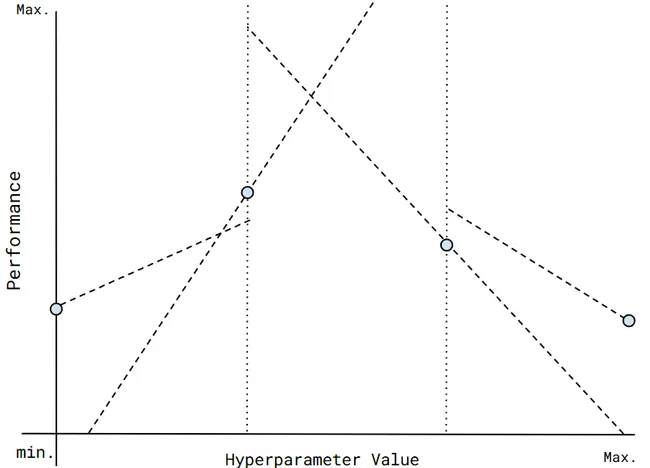
Gradient-based preprocessing concept visualization. The search space has been "split" by the fine dotted lines and gradients (dash lines) act as performance projections for each value region.
 Hyperparameter Optimization Machine Learning Model Optimization
Hyperparameter Optimization Machine Learning Model Optimization NOCTURNAL: Exploring the Dark Chemical Space
A drug discovery framework designed for drug-structure screening and lead optimization against user-defined biological targets. Paired with interactive chemical space analysis and various chemical-similarity calculations, aimed at accelerating drug candidate identification and optimization.
DataSift: Less Noise, More Performance
A feature space optimization tool for Binary Classifier efficiency optimization. Combining a thorough variance threshold optimizer and a backward-iteration importance-based feature pruner, this module is geared towards high-dimensional data where less noise == more performance.
Education
Honours Biomedical Sciences
University of Waterloo
- Graduated with Distinction